Scientists model a Coronavirus' infectious bits for the first time
Their efforts could lead to more effective SARS treatments.
A collaboration of scientists from University of Washington (UW), the Pasteur Institute and the University of Utrecht have harnessed a state-of-the-art microscope and supercomputer to model a coronavirus' infection mechanism for the first time.
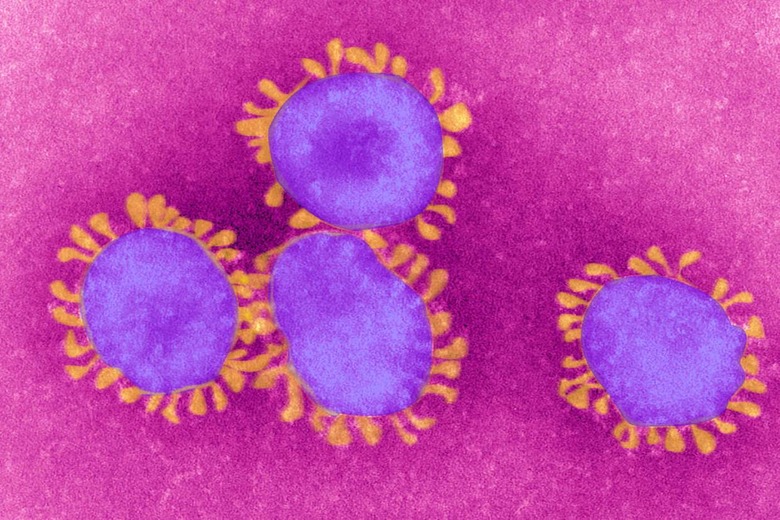
Coronaviruses are really good at infecting the respiratory systems of humans and other mammals. Once inside, these viruses can cause pneumonia (if you're lucky). The strains that become SARS and MERS have a mortality rate as high as 37 percent. Plus, there is currently no antigen for SARS or MERS, which makes them especially dangerous.
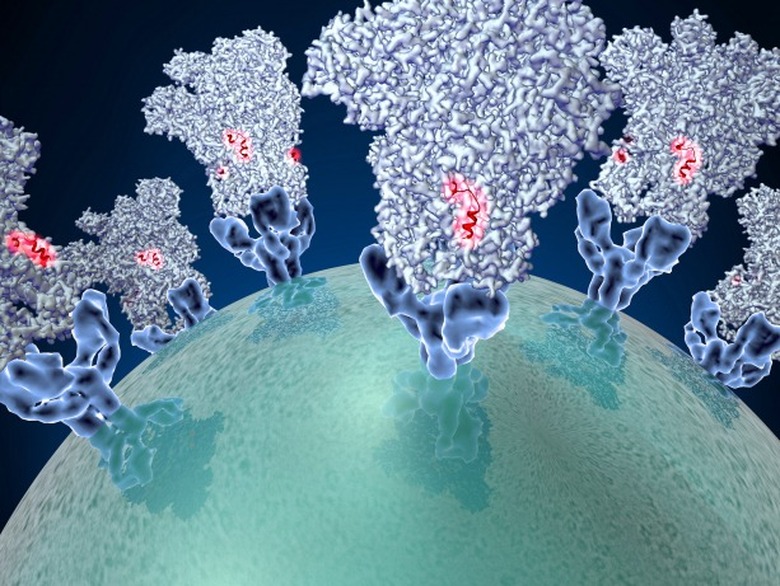
The virus is so effective because of its "transmembrane spike glycoprotein," which binds to the surfaces of other cells, allowing the virus to enter. This structure is what gives coronaviruses their spiky, crown-like shape and determines what species of animals the virus can target.
The research team leveraged a single particle cryo-electron microscopy technique to model the spike of a coronavirus that infects mice in terrific detail. The team managed a 4 angstrom resolution — about a tenth of a nanometer.
With this new analysis, the team believes they've identified a potential weaknesses in the virus' defenses. Turns out, the spike has a small peptide chain running along it. That peptide helps facilitate the virus' entry into a cell but could easily be hijacked by a treatment.
"Small molecules or protein scaffolds might eventually be designed to bind to this site," UW assistant professor of chemistry, David Veesler said in a statement, "to hinder insertion of the fusion peptide into the host cell membrane and to prevent it from undergoing changes conducive to fusion with the host cell. We hope that this might be the case, but much more work needs to be done to see if it is possible."